Search Docs by Keyword
> User Quick Start Guide
This guide will provide you with the basic information needed to get up and running on the FASRC cluster. If you’d like more detailed information, each section has a link to more full documentation
|
LOGIN USERNAME CHEATSHEET CLI/Portal/VDI/OoD – For the majority of our services you will log in with your FASRC username and password. Your username was selected at signup. You will set your password using the instructions below (step 2). Your two-factor verification code may also be required. VPN – When connecting to our VPN you will still use your FASRC username, but you will also need to specify what VPN realm you wish to connect to. Realms provide access to different environments. For instance, the FASSE secure environment uses a special realm ‘@fasse’. Unless you have been told to use a different realm, you will want to use the ‘@fasrc’ realm. Email Address – You will not use your email address to log into any FASRC service. The only FASRC system where you will need to enter your email address is when setting or resetting your FASRC account password (see step 2 below). Harvard Key – Similarly, you will not use your Harvard Key to log into FASRC services. The single exception to this is for account approvers to log in to approve new accounts. Two-Factor – Also, Harvard Key is not your two-factor code provider. FASRC has its own two-factor authentication. When logging in to FASRC systems you will sometimes be asked for this 6-digit token code. While you can use Duo to store this token, again it is not tied to Harvard Key in any way. Please see OpenAuth below (step 3). |
PREREQUISITES
1. Get a FASRC account using the account request tool.
Before you can access the cluster you need to request a Research Computing account.
See How Do I Get a Research Computing Account for instructions if you do not yet have an account.
Once you have your new FASRC account, you will need to to do two things to allow you to authenticate: Set your password and set up your OpenAuth (2FA) token.
2. Set your FASRC password
You will be unable to login until you set your password via the RC password reset link: https://portal.rc.fas.harvard.edu/p3/pwreset/
You will need to enter the same email address you used to sign up for your account and then will receive an email with a reset link (this email and link expires after 15 minutes and is for one-time use – it is never needed again). Once you’ve set your password you can use your username and password to request your OpenAuth token.
3. Set up OpenAuth for two-factor authentication (2FA)
NOTE: This is not your Harvard Key two-factor code. FASRC has its own two-factor system.
You will need to set up our OpenAuth two-factor authentication (2FA) either on your smartphone (using Google Authenticator, Duo, or similar OTP app) or by downloading our Java applet on your computer.
See the OpenAuth Guide for instructions if you have not yet set up OpenAuth.
For troubleshooting issues you might have, please see our troubleshooting page.
4. Use the FASRC VPN when connecting to storage, VDI, or other resources
Note that VPN is not required to SSH into a login node, but is required for many other tasks including VDI and mounting shares on your desktop/laptop.
5. Review our introductory training
See: Introduction to Cluster Computing Docs and Video
Accessing the Cluster and Cluster Resources
Terminal access via SSH to login.rc.fas.harvard.edu
[wpfa5s icon=”fa-exclamation-circle”] If you did not request cluster access when signing up, you will not be able to log into the cluster or login nodes as you have no home directory. You will simply be asked for your password over and over. See this doc for how to add cluster access as well as additional groups.
For command line access to the cluster, you will SSH to login.rc.fas.harvard.edu using your FASRC username, password and OpenAuth token.
[wpfa5s icon=”fa-info-circle”] See our SSH/Command Line access documentation for detailed instructions: Command line access using a terminal
Watch our Getting Started on the FASRC Cluster video
Graphical Desktop Access using VDI/OpenOnDemand
We also provide an interactive graphical desktop environment from which you can launch graphical applications as well as SSH/command line jobs. Please remember that you must be connected to our VPN to access this service.
See the following docs for more details: Virtual Desktop through Open OnDemand
File Transfer
[wpfa5s icon=”fa-info-circle”] See our Transferring Data on the Cluster page for best practices for moving data around. Our SCP page offers additional details for SCP.
There are also graphical scp tools available. The Filezilla SFTP client is available cross-platform for Mac OSX, Linux, and Windows. See our SFTP file transfer using Filezilla document for more information. Windows users who prefer SCP can download it from WinSCP.net and follow the steps from Connecting with WinSCP to connect to Cannon.
[wpfa5s icon=”fa-exclamation-circle”] If you are off-campus or behind a firewall and wish to connect to FASRC servers other than the login servers, you should first connect to the Research Computing VPN.
Determine where your files will be stored
Watch our Introduction to Cluster Computing Storage video
Users of the cluster are granted 100Gb of storage in their home directory. This volume has decent performance and is regularly backed up. For many, this is enough to get going. However, there are a number of other storage locations that are important to consider when running software on the FASRC cluster.
-
- /n/holyscratch01 – Our global scratch (environment variable $SCRATCH) is large, high performance temporary Lustre filesystem. We recommend that people use this filesystem as their primary job working area, as this area is highly optimized for cluster use. Use this for processing large files, but realize that files will be removed after 90 days and the volume is not backed up. Create your own folder inside the folder of your lab group. If that doesn’t exist, contact RCHelp.
- /scratch – Local on-node scratch. When running batch jobs (see below), /scratch is a large, very fast temporary store for files created while a tool is running. This space is on the node’s local hard drive. It is a good place for temporary files created while a tool is executing because the disks are local to the node that is performing the computation making access is very fast. However, data is only accessible from that node so you cannot directly retrieve it after calculations are finished. If you use /scratch, make moving any results off and onto another storage system part of your job.
- Lab storage – Each lab that is doing regular work on the cluster can request an initial 4Tb of group accessible storage at no charge. Like home directories, this is a good place for general storage, but it is not high performance and should not be used during I/O intensive processing. See the global scratch above.
Do NOT use your home directory or lab storage for significant computation.
This degrades performance for everyone on the cluster.
For details on different types of storage and how to obtain more, see the Cluster Storage page.
Running Jobs and Loading Software
Software Modules
An enhanced module system called Helmod is used on the cluster to control the run-time environment for individual applications. To find out what modules are available we recommend you can check the module list on the RC / Informatics portal. You can also use the module avail command, although it will be less ionformative. By itself, module avail will print out the entire list of packages. To find a specific tool, use the module spider or module-query command.
module-query MODULENAMEOnce you’ve determined what software you would like to use, load the module:
module load MODULENAMEwhere MODULENAME is the specific software you want to use. You can use module unload MODULENAME to unload a module. To see what modules you have loaded type module list.
TIP: This is very helpful information to provide when you submit help tickets.
For details on finding and using modules effectively, see Software on the cluster page.
For more details on running software and jobs on the cluster, including graphical applications, see module section of the Running Jobs page.
Run a batch job…
The cluster is managed by a batch job control system called SLURM. Tools that you want to run are embedded in a command script and the script is submitted to the job control system using an appropriate SLURM command.
For a simple example that just prints the hostname of a compute host to both standard out and standard err, create a file called hostname.slurm with the following content:
#!/bin/bash #SBATCH -c 1 # Number of cores requested #SBATCH -t 15 # Runtime in minutes #SBATCH -p serial_requeue # Partition to submit to #SBATCH --mem=100 # Memory per node in MB (see also --mem-per-cpu) #SBATCH --open-mode=append # Append when writing files #SBATCH -o hostname_%j.out # Standard out goes to this file #SBATCH -e hostname_%j.err # Standard err goes to this filehostname hostname
Then submit this job script to SLURM
sbatch hostname.slurmWhen command scripts are submitted, SLURM looks at the resources you’ve requested and waits until an acceptable compute node is available on which to run it. Once the resources are available, it runs the script as a background process (i.e. you don’t need to keep your terminal open while it is running), returning the output and error streams to the locations designated by the script.
Please note that your output and error files will be written to the directory the job starts in unless you specify the full path. So, for example, if you always want your error files to write to your home directory, you would use:
#SBATCH -e /n/home05/myusername/hostname_%j.err
Note that home05 may be different for you. You can find the full path of your home directory with the pwd command:
[jharvard@boslogin01 ~]$ pwd /n/home01/jharvard
You can monitor the progress of your job using the squeue -j JOBID command, where JOBID is the ID returned by SLURM when you submit the script. The output of this command will indicate if your job is PENDING, RUNNING, COMPLETED, FAILED, etc. If the job is completed, you can get the output from the file specified by the -o option. If there are errors, that should appear in the file specified by the -e option.
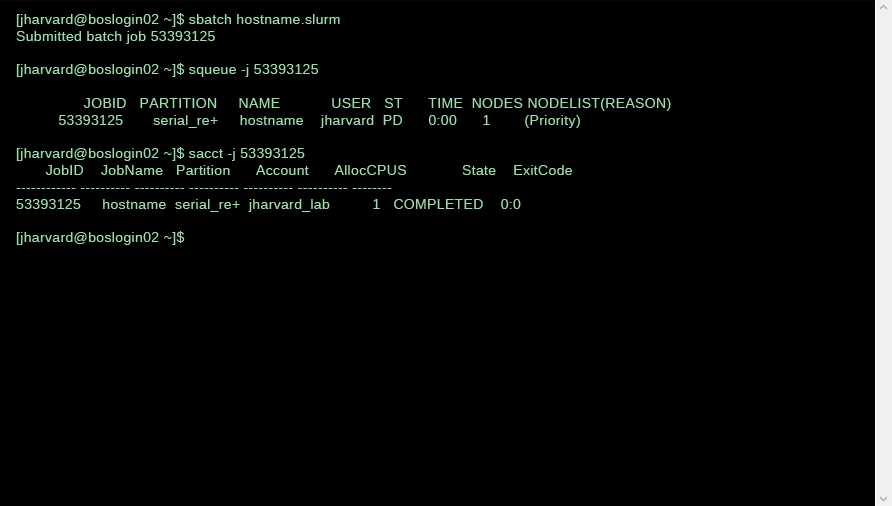
If you need to terminate a job, the scancel command can be used (JOBID is the number returned when the job is submitted).
scancel JOBIDSLURM-managed resources are divided into partitions (known as queues in other batch processing systems). Normally, you will be using the shared or serial_requeue partitions, but there are others for interactive jobs (see below), large memory jobs, etc.
For more information on the partitions on the cluster, please see the SLURM partitions page.
For more detailed information on running batch jobs, including MPI code, please see the Running Jobs page.
For a list of useful SLURM commands, please see the Convenient SLURM Commands page.
… or run an interactive job.
Batch jobs are great for long-lasting computationally intensive data processing. However, many activities like one-off scripts, graphics and visualization, and exploratory analysis do not work well in a batch system but are too resource-intensive to be done on a login node. There is a special partition on the cluster called “test” that is designed for responsive, interactive shell and graphical tool usage.
You can start an interactive session using a specific flavor of the salloc command.
salloc -p test --mem 500 -t 0-08:00salloc is like sbatch, but it runs synchronously (i.e. it does not return until the job is finished). The example starts a job on the “test” partition, an allocation of 500 MB RAM (--mem 500), and for 6 hours (-t in D-HH:MM format). It also assumes one core on one node. The final argument is the command that you want to run. In this case you’ll just get a shell prompt on a compute host. Now you can run any normal Linux commands without taking up resources on a login node. Make sure you choose a reasonable amount of memory (--mem) for your session.
For graphical tools and applications, we also offer a virtual graphical desktop through our VDI/OpenOnDemand system. See the following docs for more details: Virtual Desktop through Open OnDemand
A note on requesting memory (--mem or --mem-per-cpu)
In SLURM you must declare how much memory you are using for your job using the --mem or --mem-per-cpu command switches. By default, SLURM assumes you need 100 MB. If you don’t request enough the job can be terminated, oftentimes without very useful information (error files can show segfault, file write errors, etc. that are downstream symptoms). If you request too much, it can increase your wait time (it’s harder to allocate a lot of memory than a little), crowd out jobs for other users, and lower your fairshare.
You can view the runtime and memory usage for a past job with
sacct -j JOBID --format=JobID,JobName,ReqMem,MaxRSS,Elapsedwhere JOBID is the numeric job ID of a past job:
sacct -j 531306 --format=JobID,JobName,ReqMem,MaxRSS,ElapsedJobID JobName ReqMeM MaxRSS Elapsed
531306 sbatch 00:02:03
531306.batch batch 750000K 513564K 00:02:03
531306.0 true 916K 00:00:00
The .batch portion of the job is usually what you’re looking for, but the output may vary. This job had a maximum memory footprint of about 500MB, and took a little over two minutes to run.
Familiarize yourself with proper decorum on the cluster
The FASRC cluster is a massive system of shared resources. While much effort is made to ensure that you can do your work in relative isolation, some rules must be followed to avoid interfering with other user’s work.
The most important rule on the cluster is to avoid performing computations on the login nodes. Once you’ve logged in, you must either submit a batch processing script or start an interactive session (see below). Any significant processing (high memory requirements, long running time, etc.) that is attempted on the login nodes will be killed.
See the full list of Cluster Customs and Responsibilities.
Getting further help
If you have any trouble with running jobs on the cluster, first check the comprehensive Running Jobs page and our FAQ. Then, if your questions aren’t answered there, feel free to contact us at RCHelp. Tell us the job ID of the job in question. Also provide us with what script you ran, the error and output files, and where they’re located as well. The output of module list is helpful, too.
